Solaris 2.0
The new and improved Solaris 2.0 amplification chemistry and flexible library preparation workflows improve both speed and data quality.
Speak with a Specialist

Improved GC coverage with Solaris 2.0
Ultima integrates enhanced sequencing amplification chemistry with flexible library preparation options, including both PCR-free and indexing PCR workflows. Together, these improvements reduce GC bias and improve coverage uniformity across the genome. The data shown here were generated using the Solaris 2.0 PCR-free library workflow.
Download WGS application noteAccurate variant calling across genome
Variant calling performance was assessed in comparison to reference GIAB truth sets, using the full NIST HCR v4.2.1 for each reference sample unless otherwise noted. The overall average concordance of single nucleotide polymorphism (SNP) variant calls over the dataset was F1 = 99.79%, and for Indel variant calls (excluding homopolymers ≥ 12) was F1= 99.06%. Variant calling in coding regions returned SNP F1 = 99.60% and Indel F1 = 98.84%.


Clinically relevant gene coverage
The improvement in high-GC coverage translates into more consistent depth across GC-rich, clinically relevant loci that are traditionally challenging for short-read sequencing workflows. By reducing dropout and uneven amplification in these regions, Solaris 2.0 enables more reliable variant detection, improved confidence in calling across promoter regions and GC-dense exons, and stronger performance in genes associated with inherited disease and oncology applications.
Download germline WGS application noteFaster workflow and fewer hands-on steps
Solaris 2.0 provides a simplified 7-hour amplification workflow with hands-on time of only 10 minutes with one touch point. When using UG200 Series sequencers, libraries can be amplified and sequenced in 13 to 19 hours total depending on the sequencing read length.
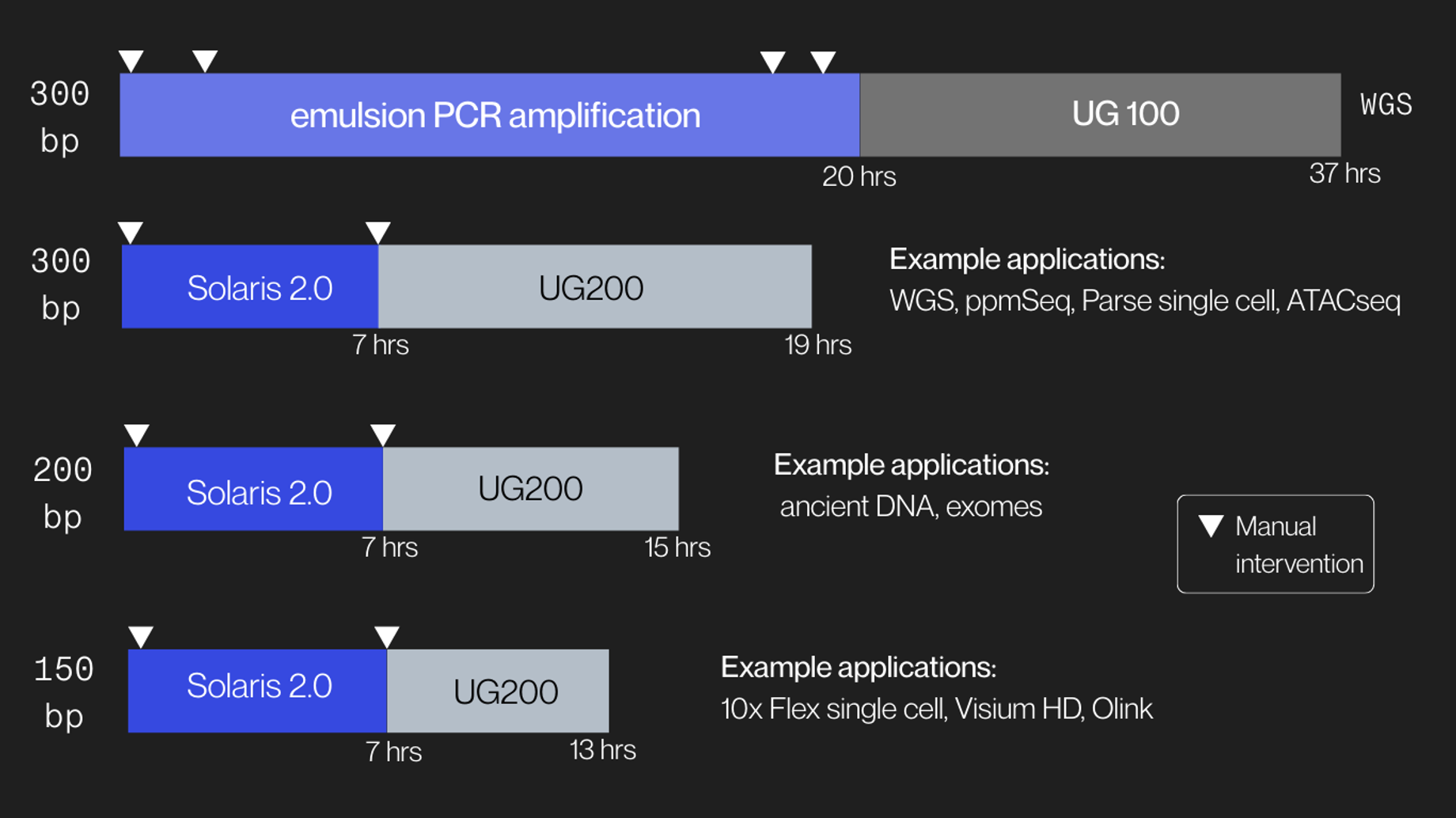
Solaris 2.0 Free libary prep
The Solaris 2.0 Free workflow offers PCR-free library prep compatible with leading library prep kits and accommodates a range of sample types and input amounts.
Highlighted Solaris 2.0 Free applications:
Germline whole genome sequencing
Somatic whole genome sequencing
ppmSeq for tumor/normal pairs
ppmSeq for liquid biopsy and MRD

Solaris 2.0 Flex libary prep
The Solaris 2.0 Flex workflow allows for seamless adaptation of partial or full-length existing libraries generated from third-party library kits through a simple PCR.
Highlighted Solaris 2.0 Flex applications:
Single-cell sequencing
Methylation sequencing
RNA sequencing
Proteomics

Solaris 2.0 PCR-Free Adapters available from Ultima Genomics
Quantify Solaris 2.0 Free Libraries: NEBNext Library Quant Kit
Solaris 2.0 Flex Indexing PCR Primers available from Ultima Genomics
Solaris 2.0 library prep workflow highlights
| Workflow Name | Adapter & Primer Names | Sample indexing | DNA Type | DNA Input Range |
|---|---|---|---|---|
| Solaris 2.0 Free | Solaris 2.0 PCR-Free Adapters | 192 sample indices | Genomic DNA (gDNA) | 100 - 250 ng* |
| Solaris 2.0 Flex | Solaris 2.0 Indexing and UG-Universal Primers | 384 sample indices | Native or existing library | 10 ng* |
*DNA input amounts verified by Ultima Genomics.
A growing ecosystem of library prep partners
Start saving now.
Run your next project on an Ultima sequencer.

Certified Service Providers
Submit samples and start generating data on an Ultima sequencer. Access cost-effective and scalable sequencing with a growing network of global service providers rigorously vetted for their technical expertise and adherence to best practices.
Sequence with a CSP todayTechnology Access Program
Our Technology Access Program offers accessible entry to Ultima sequencing through our in-house applications lab. Sequence on our high-capacity fleet of Ultima sequencers and receive support at every step, from consultation through data analysis.
Discover our TAP offeringsBring Ultima into your lab
Interested in exploring how your lab can benefit from high-throughput, cost-effective sequencing? Learn how our novel use of open wafers and built-in automation is continuously driving the cost of sequencing down to accelerate omics at scale.
Speak with a Sales Specialist